Sepsis Watch
GA Tech College of Computing / Emory Department of Biomedical Informatics
Sepsis is the body’s overpowering reaction to infection which can lead to organ damage, organ failure, and death. Approximately 27 million people in the world develop sepsis each year, and, in Georgia alone, the mortality rate was 85 deaths per 100,000 persons. Total spending in US hospitals in 2011 was 20.3 billion, which was 5.2% of hospital costs. For each hour that treatment (antibiotics) of sepsis is delayed, mortality increases by about 8%.
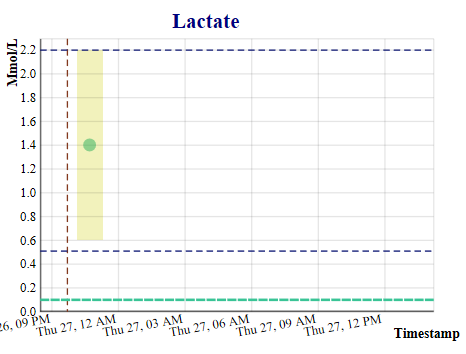
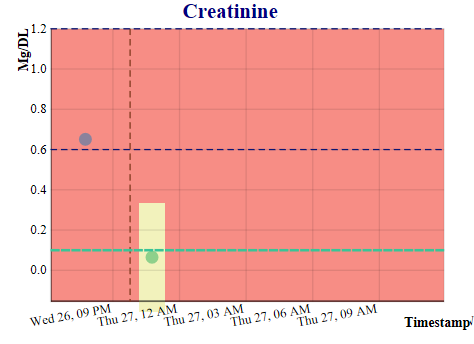
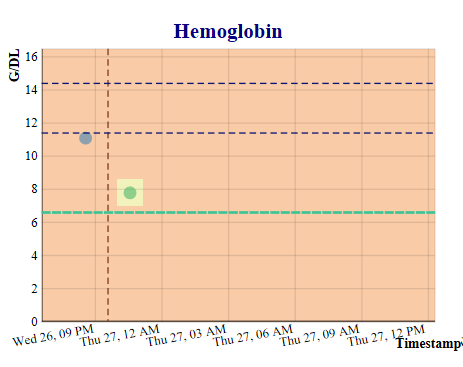
This application was developed as part of a project for a health informatics class at Georgia Institute of Technology, is for demonstration purposes only, and is not yet approved for clinical use. The purpose of this application is to predict the values of six labs which are advance indicators of sepsis, so that the progress of sepsis can be prevented by proactive treatment. Using an extract of the MIMIC III critical care database (100 patients), an openly available dataset developed by the MIT Lab for Computational Physiology, which includes laboratory tests, vital signs, medications, timestamps, and more, we developed a Gaussian process prediction model for white blood counts and five other relevant lab tests, including hemoglobin, bilirubin, platelets, creatinine, and lactates. The application allows the user to select a physician, and then a patient treated by that physician, launching the display of six screens with both the actual and predicted lab values for that patient.
The prediction models will be implemented with a FHIR server providing electronic health record actual lab values from the electronic health record and a second server providing streaming vitals information from the Philips Bedmaster, so that the predictions can be updated in real time, each time new data is received by the application. The prediction algorithm is executed when new data is received from either source. When a lab prediction is in the critical range, a mobile message is sent to the physician for evaluation and possible action. The physician can then make a decision on whether additional labs, antibiotics, or other treatment is needed for the patient. This work will also have implications for the computational phenotyping of sepsis, savings on lab tests, and provision of general trending information of the patient's health to the physician.
Designed for
Application Type
FHIR Compatibility
Categories
Specialties
Licensing & Pricing
OS Support
Report Listing